smoothdata
Smoothing of noisy data.
| Library |
|
Syntax
Function call
-
B = smoothdata(A)— smooths out the elementsAusing a moving average. Functionsmoothdatadetermines the size of the sliding window based on the elementsA. The window is moved along the length of the vector, calculating the average value for the elements of each window. IfA— the matrix, then the functionsmoothdatacalculates the moving average for each columnA.
-
B = smoothdata(_, Name,Value)— uses additional smoothing parameters specified by one or more arguments of the typeName,Value.
Arguments
Input arguments
# A — input data
+
vector | the matrix
Details
Input data specified as a vector or matrix.
| Типы данных |
|
| Support for complex numbers |
Yes |
# dim — the measurement for which the operation is performed
+
scalar
Details
The dimension that the operation is performed on, specified as a positive integer scalar. If no dimension is specified, then the first dimension of the array is used by default, the size of which is not equal to 1.
Consider the input matrix A size on :
# method — smoothing method
+
"movmean" (by default) | "movmedian" | "gaussian" | "lowess" | "loess" | "rlowess" | "rloess" | "sgolay"
Details
The smoothing method specified by one of the following values:
-
"movmean"— average for each windowA. This method is useful for reducing periodic trends in data. -
"movmedian"— median for each windowA. This method is useful for reducing periodic trends in data when outliers are present. -
"gaussian"— Gaussian weighted average for each windowA. -
"lowess"— linear regression for each windowA. This method can be computationally expensive, but it leads to fewer gaps. -
"loess"— quadratic regression for each windowA. This method is more computationally expensive than"lowess". -
"rlowess"— stable linear regression for each windowA. This method is a more computationally expensive version of the method."lowess", but more resistant to emissions. -
"rloess"— robust quadratic regression for each windowA. This method is a more computationally expensive version of the method."lowess", but more resistant to emissions. -
"sgolay"— Savitsky filter — Golay, which smooths the data according to a quadratic polynomial selected for each windowA. This method may be more effective than other methods when data is changing rapidly.
# window — window size
+
scalar | two-element vector
Details
The window size specified as a positive integer scalar or a two-element vector of non-negative integers. Function smoothdata defines the window relative to the sampling points.
-
If
window— this is a positive integer scalar, then the window has lengthwindowand it is centered relative to the current element. -
If
windowis a two—element vector of non-negative integers[b f], that window contains the current element,bthe preceding elements andfsubsequent elements.
For more information about the window position, see Sliding window size.
# nanflag — condition for missing value
+
"omitmissing" (by default) | "omitnan" | "includemissing" | "includenan"
Details
The condition for missing a value set by one of the following values:
-
"omitmissing"or"omitnan"— ignore the valuesNaNinAwhen smoothing. If all the elements are in the window —NaN, then the corresponding elements inBalsoNaN. Values"omitmissing"and"omitnan"they behave the same way. -
"includemissing"or"includenan"— enable valuesNaNinAwhen smoothing. If any element is in the window —NaN, then the corresponding elements inBalsoNaN. Values"includemissing"and"includenan"they behave the same way.
Name-value input arguments
Specify optional argument pairs in the format Name, Value, where Name — the name of the argument, and Value — the appropriate value. Name-value arguments should be placed after other arguments, but the order of the pairs does not matter.
Use commas to separate the name and value, and Name put it in quotation marks.
Example: B = smoothdata([4.2, 3.8, 4.3, 5], "SmoothingFactor", 0.5).
# SmoothingFactor — window size factor
+
scalar
Details
The window size coefficient, set as a scalar in the range from 0 before 1. As a rule, the value SmoothingFactor adjusts the anti-aliasing level by scaling the window size that the function smoothdata determines based on the values in A. Values close to 0, result in a smaller window size, which results in less anti-aliasing. Values close to 1, lead to an increase in the window size, which leads to more anti-aliasing. In some cases, depending on the values used smoothdata to determine the window size, the value of the argument is SmoothingFactor it may not have a significant effect on the window size.
The default value of the argument is SmoothingFactor equally 0.25. Point out SmoothingFactor it is allowed only if not specified window.
# Degree — the degree of the Savitsky filter — Goleya
+
scalar
Details
Degree of the Savitsky filter — A golem defined as a non-negative integer scalar. This name-value argument can only be specified if the smoothing method is selected. "sgolay". The value of the argument Degree corresponds to the degree of the polynomial in the Savitsky filter — A goal that approximates the data in each window. By default, it is equal to 2.
The value of the argument Degree it should be smaller than the window size for uniform sampling points. For uneven sample points, the value should be less than the maximum number of points in any window.
Output arguments
# winsize — window size
+
scalar | two-element vector
Details
The window size returned as a positive integer scalar or a two-element vector of non-negative integers.
Examples
Smoothing data using a moving average
Details
Let’s create a vector containing noisy data, and smooth the data using a moving average. Let’s plot the original and smoothed data.
import EngeeDSP.Functions: smoothdata
using Plots, Random
Random.seed!(0)
x = 1:100
A = cos.(2π * 0.05 * x .+ 2π * rand()) .+ 0.5 * randn(100)
B = smoothdata(A)
plot(x, A, label="Input Data")
plot!(x, B, label="Smoothed Data")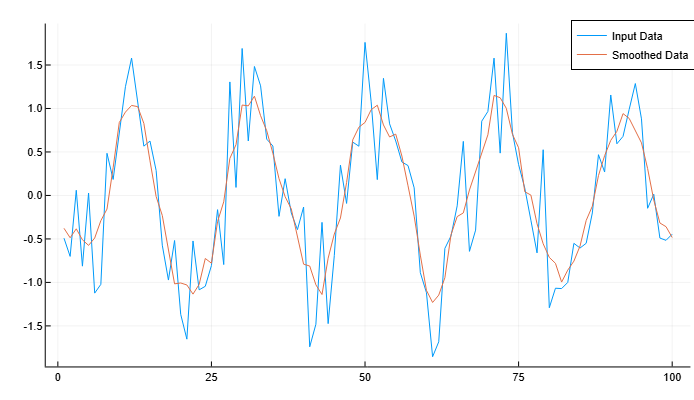
The noisy data matrix
Details
Let’s create a matrix, the rows of which represent three noisy signals. Let’s smooth out these three signals using a moving average and plot the smoothed data.
import EngeeDSP.Functions: smoothdata
using Plots, Random
Random.seed!(0)
x = 1:100
s1 = cos.(2π * 0.03 * x .+ 2π * rand(rng)) .+ 0.5 * randn(rng, 100)
s2 = cos.(2π * 0.04 * x .+ 2π * rand(rng)) .+ 0.4 * randn(rng, 100) .+ 5
s3 = cos.(2π * 0.05 * x .+ 2π * rand(rng)) .+ 0.3 * randn(rng, 100) .- 5
A = [s1'; s2'; s3']
B = smoothdata(A, 2)[1]
plot(x, B[1, :], label="s1")
plot!(x, B[2, :], label="s2")
plot!(x, B[3, :], label="s3")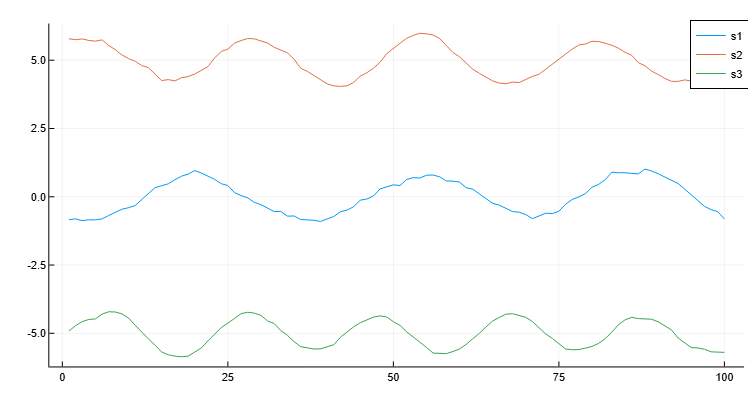
Gaussian filter
Details
Let’s smooth out the vector of noisy data using a Gaussian-weighted moving average filter. Let’s display the size of the window used by the filter.
import EngeeDSP.Functions: smoothdata
using Plots, Random
Random.seed!(0)
x = 1:100
A = cos.(2π * 0.05 * x .+ 2π * rand(rng)) .+ 0.5 * randn(rng, 100)
B, winsize = smoothdata(A, "gaussian")
winsize4.0Flatten the original data using a larger window containing 20 elements. Let’s plot the smoothed data for windows of both sizes.
C = smoothdata(A, "gaussian", 20)[1]
plot(x, B, label="Small Window", linewidth=2)
plot!(x, C, label="Large Window", linewidth=2)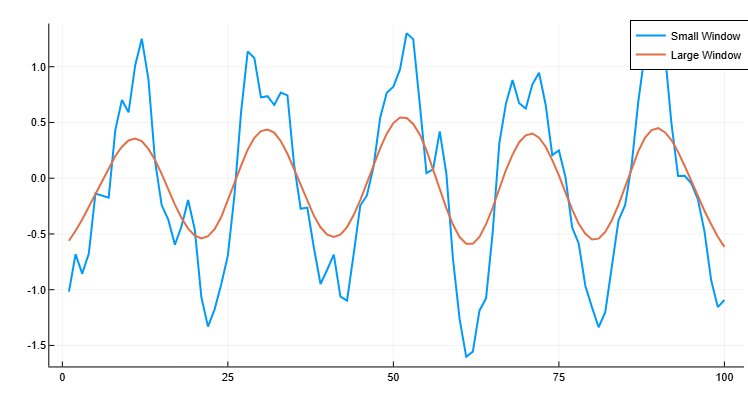
Smoothing data with missing values
Details
Create a noisy vector containing the values NaN, and smooth the data, ignoring the values NaN.
import EngeeDSP.Functions: smoothdata
A = [NaN randn(1,48) NaN randn(1,49) NaN]
B, w = smoothdata(A)([NaN 0.5580616078303169 … 1.0570367844487276 2.3203045641188864], 2.0)Smoothing out the data, including the values NaN. The average value in a window containing at least one value NaN, equal to NaN.
C, w = smoothdata(A, "includenan")([NaN NaN … 1.0570367844487276 NaN], 2.0)Let’s plot the smoothed vectors B and C.
plot(1:100, B', shape=:circle, label="Ignore Missing")
plot!(1:100, C', shape=:xcross, label="Include Missing")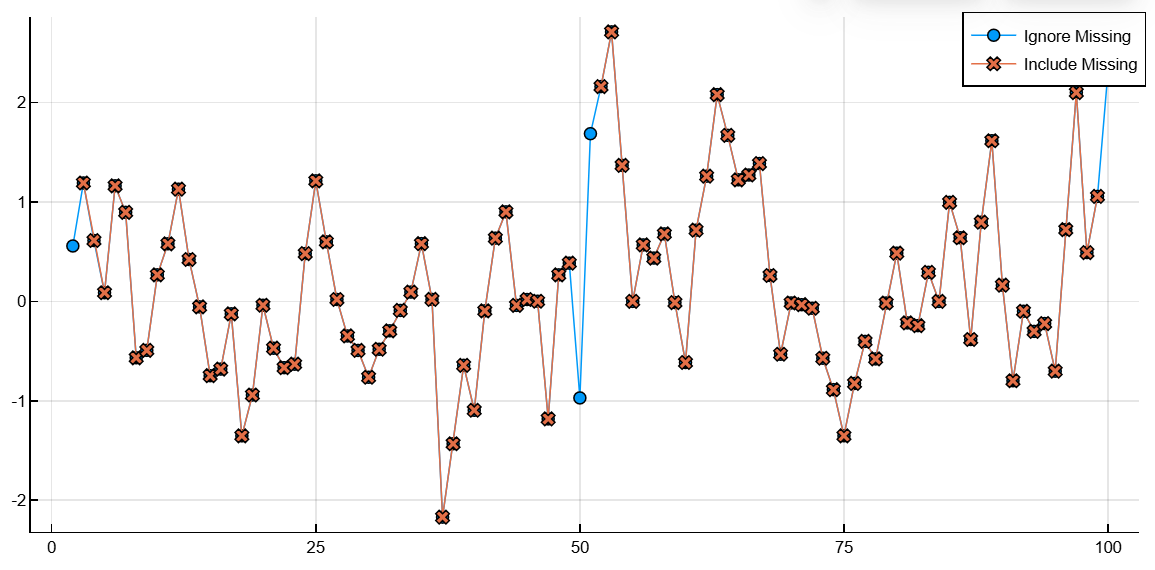
Additional Info
Sliding window size
Details
This table shows the position of the window according to the vector of evenly distributed sample points by default. [1 2 3 4 5 6 7].
| Description | Window size and position | Sample points in the window | Scheme |
|---|---|---|---|
For a scalar window size, the front edge of the window is included and the back edge of the window is excluded. |
|
|
|
|
|
||
To determine the size of a vector window, the leading and trailing edges are taken into account. |
|
|
|
For sampling points located near the endpoints of the input data, these sliding statistics smoothing methods truncate the window so that it starts at the first sampling point or ends at the last:
|
|
|
|
For sampling points located near the endpoints of the input data, these local regression smoothing methods shift the window so that it includes the first or last sampling point.:
|
|
|
Algorithms
If the window size is not specified for the smoothing method, then the function smoothdata calculates the default window size based on heuristics. For the smoothing coefficient The heuristic estimates the size of the moving average window, which weakens approximately percent of the input data energy.






